Ministry of Science & Technology
Altered understanding of bacterial transcription mechanism can pave new path to combat tuberculosis
Posted On:
02 APR 2026 3:31PM by PIB Delhi
A team of scientists has uncovered a fundamental flaw in a long-standing model of how bacteria control gene expression that can form the base for innovative strategies to combat tuberculosis and other bacterial infections.
Tuberculosis is one of the world’s deadliest infectious diseases, and drug-resistant strains pose an increasing global threat to cure of the disease. M. tuberculosis (TB) bacteria survive inside the human host by precisely regulating gene expression under extreme stress conditions.
For years, scientists believed that a protein called σ factor binds RNA polymerase, initiates bacterial transcription and is then released once the enzyme begins elongating RNA. This process, known as the σ-cycle, was assumed to be universal across bacteria, including TB bacteria.
A new study from Bose Institute, Kolkata, an autonomous institute of the Department of Science and Technology (DST), overturns this assumption.
The Researchers Dr Jayanta Mukhopadhyay and Dr N Hazra found that while some σ factors in M. tuberculosis dissociate from RNA polymerase during transcription, others remain firmly attached throughout the process.
Their research, published in the international journal Nucleic Acids Research, reveals that a mechanism taught for decades in molecular biology textbooks—the so-called “universal σ-cycle”, does not apply to all bacteria or all regulatory proteins.
The study focuses on tuberculosis (TB) causing bacteria, Mycobacterium tuberculosis, and shows that different σ (sigma) factors, proteins that guide RNA polymerase to specific genes, behave in strikingly different ways during transcription, the first step of gene expression.
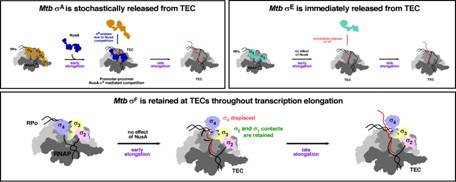
Fig: Differential Association of Different σ Factors with RNA Polymerase of Mycobacterium Tuberculosis during Transcription Elongation
Using a combination of advanced biochemical and cellular techniques, the researchers examined three distinct σ factors. These include σA, the primary housekeeping σ factor, σE, a stress-responsive σ factor and σF linked to stress survival and adaptation.
They discovered that σA and σE are released from RNA polymerase during transcription elongation, either immediately or gradually. In contrast, σF remains stably associated with the enzyme, even as transcription proceeds. This study, published in the journal Nucleic Acids, shows that bacteria do not rely on a single universal mechanism. They used multiple, finely tuned strategies to regulate gene expression.
The discovery that σF remains bound to RNA polymerase suggests a so far unknown mechanism by which the bacterium ensures sustained expression of stress-response genes, an insight with important implications for TB biology.
By showing that σ–RNA polymerase interactions vary depending on σ-factor architecture, the study highlights new, highly specific targets for antimicrobial development. Instead of targeting enzyme active sites—where resistance often emerges—future drugs could disrupt critical protein–protein interactions essential for bacterial survival.
The team used in vitro transcription assays, fluorescence-based measurements, high-resolution protein interaction studies and in vivo validation using chromatin immunoprecipitation, followed by quantitative PCR. As antibiotic resistance continues to rise, insights like these, rooted in basic molecular understanding can advance fundamental science and may prove crucial in shaping the next generation of antimicrobial therapies.
*****
NKR/FT
(Release ID: 2248321)
Visitor Counter : 87